Omicsmaster
点击蓝字 关注我们

深度定制 让研究突破预期
条形图在科研数据展示中使用频率非常高,样式也丰富多样!下面主要为大家分享10种好看的特色条形图绘制方法,使用工具主要包含ggplot2、Origin、DeepSeek、OmicShare tools等,希望对大家有帮助。
#读取数据;
dt <- read.csv("test.csv",header = T)
#载入rstatix包;
library(rstatix)
library(dplyr)
#按分组统计信息;
stats_summary <- dt %>% df_group_by(Group) %>% get_summary_stats()
#对数据进行正态性检验(Shapiro-Wilk Normality Test);
dt %>% shapiro_test(vars = "Expressions")
# 使用两种方法进行方差齐性检验;
#Levene’s test for homogeneity of variance;
dt$Group <- as.factor(dt$Group)
dt %>% levene_test(Expressions ~ Group)
#Bartlett test of homogeneity of variances;
bartlett.test(Expressions~Group,data = dt)从两种方法的统计结果来看(p-value 分别为0.1629 和0.278),方差是齐性的。
#使用rstatix包进行方差分析;
dt %>% anova_test(Expressions~Group)
#使用基础函数进行方差分析,整体来看差异显著;
oneway<-aov(Expressions~Group,data = dt)
anova(oneway)
#使用Fisher LSD法进行均值比较;
#LSD法(Fisher’s Least Significant Difference);
#LSD法检验微小的差异,比较方便的是直接得出显著性字母标记,不需人工标记;
#install.packages("agricolae")
library("agricolae")
out <- LSD.test(oneway,"Group",p.adj="bonferroni")
# 整理用于作图的数据框
rowname<-row.names(out$means)
mean<-out$means[,1]
sd<-out$means[,2]
marker<-out$groups$groups
plotdata<-data.frame(rowname,mean,sd,marker)
plotdata
#ggplot2 绘制带显著性标记的柱状图
library("ggplot2")
p1<-ggplot(plotdata,aes(x=factor(rowname),y=mean))+geom_bar(position =position_dodge(0),fill="orange",width = 0.52,stat = "identity")
p2<-p1+geom_errorbar(aes(ymin=mean-sd,ymax=mean+sd),position=position_dodge(0.6),width=0.25)
p3<-p2+geom_text(aes(x=factor(rowname),y=mean+sd+0.1,label=marker),size=4,position = position_dodge(0.6))
p4<-p3+xlab("")+ylab("Lesion diameter (mm)")
p5<-p4+scale_y_continuous(limits = c(0, 4),expand=expansion(add = c(0, 0)))
#更改y轴显示范围,这里的expand默认为TRUE。
mytheme<-theme_classic()+theme(axis.title = element_text(size = 12),
axis.text = element_text(size=12),
panel.grid.major = element_line(color = "white"),
panel.grid.minor = element_line(colour = "white"),
axis.text.x = element_text(size = 12,angle=45,vjust=0.7,hjust=0.8,color = "black"),
axis.text.y = element_text(size = 12,color = "black"),
legend.text = element_text(size = 12),legend.title = element_blank(),
plot.margin = unit(c(0.5,0.5,0.5,0.5), "cm"))
p5+mytheme
#读入本地数据;
df <- read.csv("testdata1.csv",header = T)
#载入dplyr包;
library(dplyr)
#转成tibble格式;
dt <- as_tibble(df)
#载入gghalves包和ggplot2包;
library(ggplot2)
library(gghalves)
#自定义图表主题,对图表做精细调整;
top.mar=0.2
right.mar=0.2
bottom.mar=0.2
left.mar=0.2
#自定义绘图主题1;
mytheme1 <- theme(panel.background = element_blank(),
axis.ticks.length=unit(1.6,"mm"),
plot.margin=unit(x=c(top.mar,right.mar,bottom.mar,left.mar),
units="inches"))
#自定义绘图主题2;
# 隐藏纵轴,并对字体样式、坐标轴的粗细、颜色、刻度长度进行限定;
mytheme2<-theme_classic()+
theme(text=element_text(family = "sans",colour ="gray30",size = 12),
axis.line = element_line(linewidth = 0.6,colour = "gray30"),
axis.ticks = element_line(linewidth = 0.6,colour = "gray30"),
axis.ticks.length = unit(1.5,units = "mm"),
plot.margin=unit(x=c(top.mar,right.mar,bottom.mar,left.mar),
units="inches"))
# 无需提前计算均值和标准差,直接绘制柱状图;
p1 <- ggplot(data=dt,mapping=aes(x=Sample,y=Linalool_content,
color=Sample,fill=Sample))+
stat_summary(fun = mean,geom="col",alpha=0.6,
width = 0.4,position = "identity",show.legend=F)
# 添加误差线;
# 这里的mult即multiplier(乘数,乘子),这里将误差线缩小到原本的一半;
p2 <- p1+stat_summary(fun.data=mean_sdl, fun.args = list(mult=1),
geom="errorbar",width=0.3,position = "dodge",show.legend=F)
# 在柱状图右侧添加云雨图;
p3<-p2+geom_half_violin(show.legend=F,nudge=0.3,scale="width",width = 0.3,
side='r',adjust=0.8,trim=F,color=NA,alpha=0.4)+
geom_half_point_panel(show.legend=F,nudge=0.6,
side = "r", shape=21, size=2.5, color="white")
# 使用position_nudge函数继续调整散点偏移位置;
p3<-p2+geom_half_violin(position=position_nudge(x=0.4,y=0),
show.legend=F,scale="width",width = 0.3,
side='r',adjust=0.8,trim=F,color=NA,alpha=0.4)+
geom_half_point_panel(position=position_nudge(x=0.1,y=0),
show.legend=F,side = "r", shape=21,
size=3, color="white")
# 自定义颜色;
mycolor <- c("#6ca3d4","#bf95c1","#61c1bf","#a5d395","#edd283","#e1807e")
mycolor <- c("#0077c1","#00a99e","#6bc72b","#ff5a20","#ff1620","#752995")
# 修改坐标轴样式;
p4 <- p3 + scale_colour_manual(values=alpha(mycolor,1))+
scale_fill_manual(values=alpha(mycolor,0.4))+
scale_x_discrete( name = "") +
scale_y_continuous(name = "Linalool content",
expand = expansion(add = c(0.1,0.5)))
# 应用主题样式1;
p4+mytheme1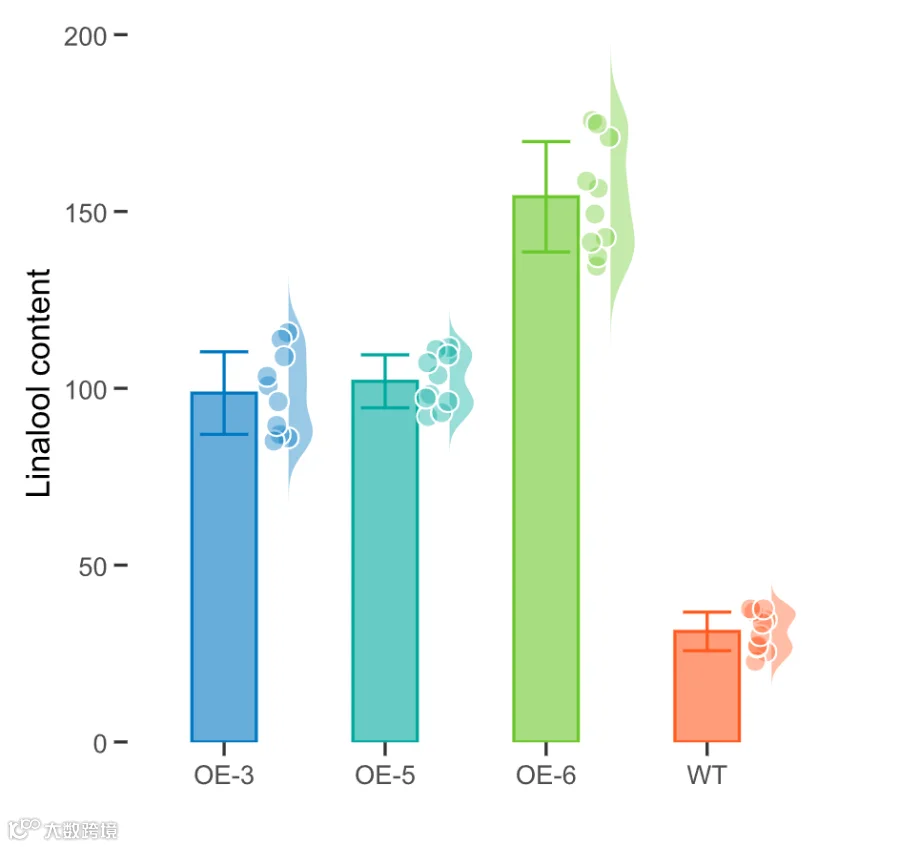
# 应用主题样式2;
p4+mytheme2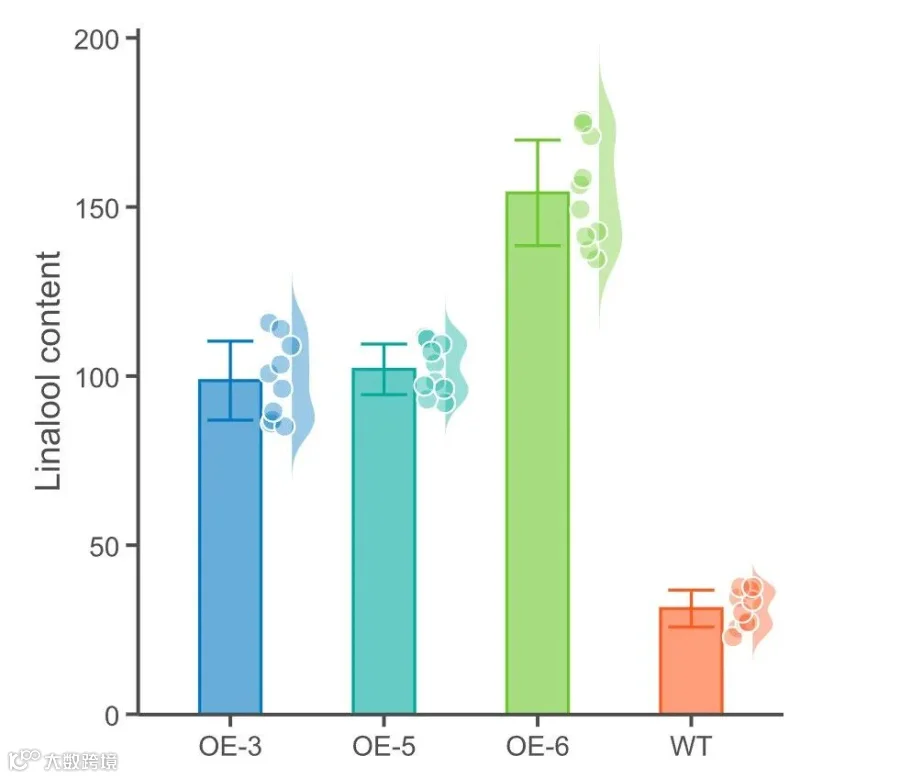
# 读取差异分析表格;
exp <- read.csv("patient-control_diff_new.csv")
# 预览数据;
head(exp)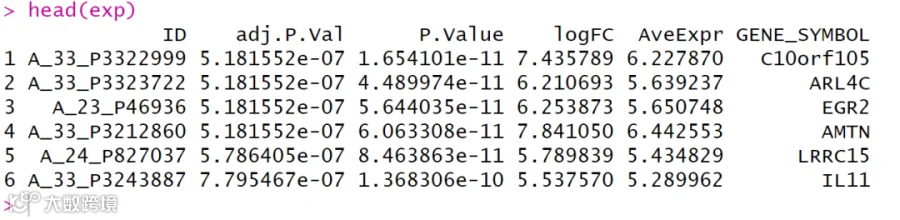
# 去掉没有symbol号的探针记录;
expr1 <- exp[exp$GENE_SYMBOL!='',]
# 查看筛选后的数据量;
dim(expr1)
#[1] 29833 6
library(dplyr)
# 根据logFC对表格进行降序排列;
expr2 <- arrange(expr1,desc(logFC))
# 根据Gene symbol列移除包含重复的行,.keep_all保留所有列;
expr3<- expr2 %>% distinct(GENE_SYMBOL, .keep_all = TRUE)
# 查看表格的行列数;
dim(expr3)
#[1] 21754 6
# 转成tibble便于后续使用,去掉不需要的列;
dt <- as_tibble(expr3[c(6,4,3)])
# 对p值取对数;
dt$log10PValue <- -log10(dt$P.Value)
# 生成显著上下调数据的分组标签;
dt$group <- case_when(dt$logFC > 1 & dt$P.Value < 0.05 ~ "Up",
dt$logFC < -1 & dt$P.Value < 0.05 ~ "Down",
abs(dt$logFC) <= 1 ~ "None",
dt$P.Value >= 0.05 ~ "None")
# 分别获取表达差异最显著的10个上下调基因;
up <- filter(dt,group=="Up") %>% top_n(10,logFC)
down <- filter(dt,group=="Down") %>% top_n(10,abs(logFC))
# 合并数据框;
df <- rbind.data.frame(up,down)
# 指定绘图顺序(从下到上),将group列转成因子型;
up$GENE_SYMBOL <- factor(up$GENE_SYMBOL,levels = rev(up$GENE_SYMBOL),ordered = T)
down$GENE_SYMBOL <- factor(down$GENE_SYMBOL,levels = rev(down$GENE_SYMBOL),ordered = T)
df$GENE_SYMBOL <- factor(df$GENE_SYMBOL,levels = rev(df$GENE_SYMBOL),ordered = T)
df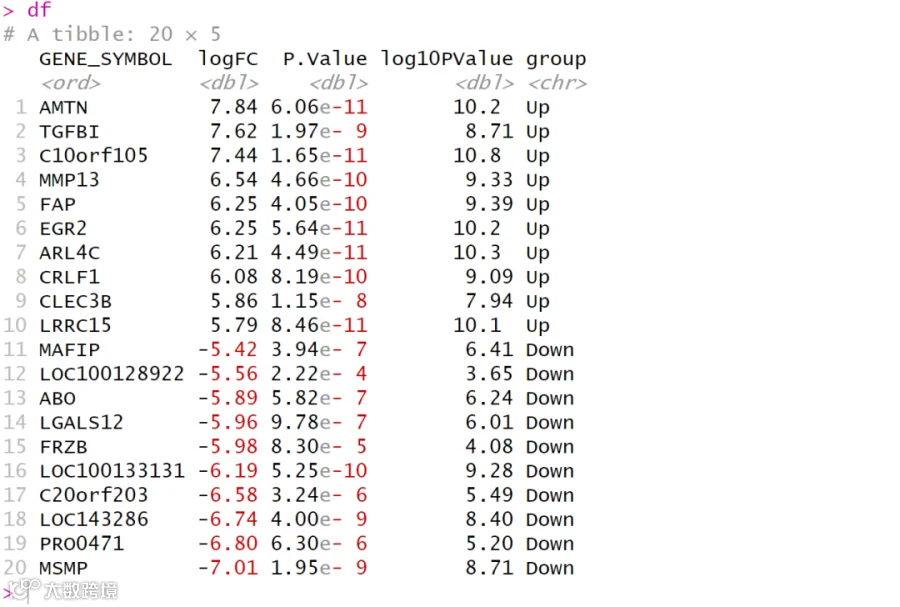
# 自定义图表主题1,对图表做精细调整;
mytheme1 <- theme_classic() +
theme(axis.text = element_blank(),
axis.ticks = element_blank(),
axis.line = element_blank(),
axis.title = element_blank(),
panel.border = element_blank(),
legend.title.position = "left",
legend.key.height=unit(0.5, "cm"),,
legend.key.width=unit(0.4, "cm"),,
legend.title = element_text(size = 9,hjust = 0.5, angle = 90),
plot.margin=unit(x=c(0.5,0.2,0,0.2),units="inches"))
# 绘制上调基因部分的图表;
f1 <- ggplot(df, aes(x = logFC,y = GENE_SYMBOL,fill = log10PValue)) +
geom_col(data=up,color = "white",width=0.9,linewidth=0.7) +
labs(x="log2FC",y="",fill="-log10 Pvalue")+
geom_text(data=up,aes(label = GENE_SYMBOL,x = lx[1:10],color = group,
hjust = ifelse(logFC < 0, 0, 1)),
show.legend = F,size = 3,fontface = "bold")+
scale_color_manual(values = mycolor[2])+
scale_fill_gradient(low = "#ffffff",high = mycolor[2],
limits = c(0,12),breaks = c(0,3,6,9,12))+
scale_x_continuous(breaks = seq(-8, 8, by = 2),
limits = c(-8, 8),expand = c(0,0)) +
geom_text(data=up,aes(label = round(logFC,2),
hjust = ifelse(logFC < 0, -0.2, 1.2),vjust = 0.5),
color="vk9pt" style="color: #576B95 !important;" data-topic="1">#ffffff",size = 3,fontface = "bold")+
mytheme1
# 自定义图表主题2,对图表做精细调整;
mytheme2 <- theme_classic() +
theme(axis.text.y = element_blank(),
axis.ticks.y = element_blank(),
axis.line.y = element_blank(),
panel.border = element_blank(),
legend.title.position = "left",
legend.key.height=unit(0.5, "cm"),,
legend.key.width=unit(0.4, "cm"),,
legend.title = element_text(size = 9,hjust = 0.5, angle = 90),
axis.line = element_line(linewidth = 0.5,lineend = "square"),
plot.margin=unit(x=c(0,0.2,0.1,0.2),units="inches"))
# 绘制下调基因部分的图表;
f2 <- ggplot(df, aes(x = logFC,y = GENE_SYMBOL,fill = log10PValue)) +
geom_col(data=down,color = "white",width=0.9,linewidth=0.7) +
labs(x="log2FC",y="",fill="-log10 Pvalue")+
geom_text(data=down,aes(label = GENE_SYMBOL,x = lx[11:20],color = group,
hjust = ifelse(logFC < 0, 0, 1)),show.legend = F,
size = 3,fontface = "bold")+
scale_color_manual(values = mycolor[1])+
scale_fill_gradient(low = "#ffffff",high = mycolor[1],
limits = c(0,12),breaks = c(0,3,6,9,12))+
scale_x_continuous(breaks = seq(-8, 8, by = 2),
limits = c(-8, 8),expand = c(0,0)) +
geom_text(data=down,aes(label = round(logFC,2),
hjust = ifelse(logFC < 0, -0.2, 1.2),vjust = 0.5),
color="#ffffff",size = 3,fontface = "bold")+
mytheme2
# 载入cowplot包进行上下拼图;;
library(cowplot)
plot_grid(f1, f2, ncol = 1, align = "v")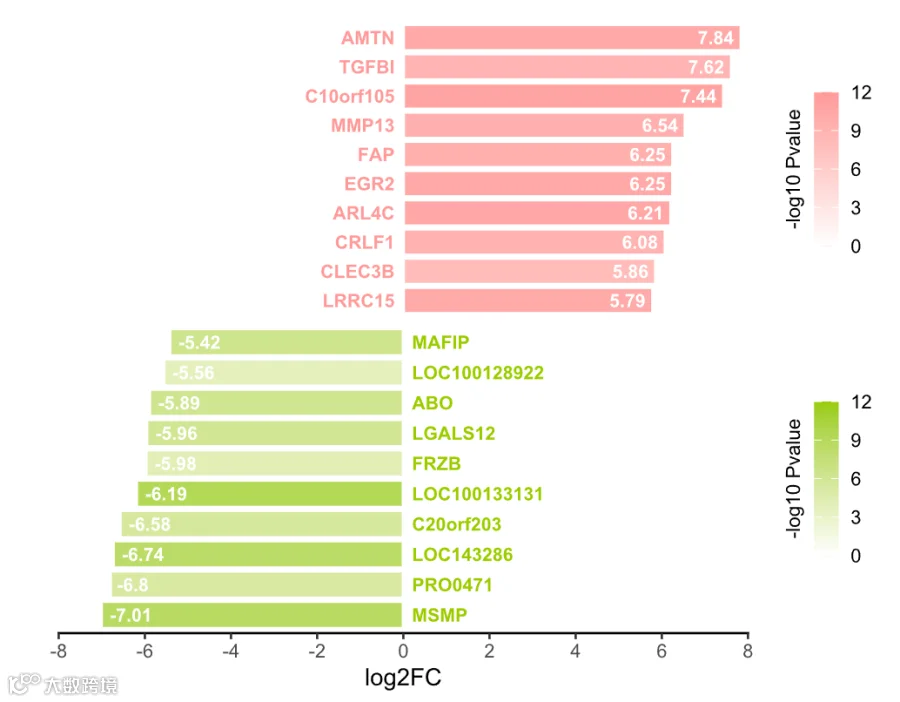
# 读入绘图数据;
dt <- read.table("out.Level2.xls",header = T,sep="\t")
head(dt,10)
# 载入所需R包;
library(dplyr)
# 按分组对数据进行降序排列;
df <- dt %>% arrange(Class,desc(number_of_gene))
# 转成因子,防止绘图时重新排序;
df$GO_Term <- factor(df$GO_Term,levels=df$GO_Term,ordered=TRUE)
# 载入ggplot2绘图包;
library(ggplot2)
# 绘制柱状图,隐藏横轴标签、标题、刻度、线条等;
p1 <- ggplot(df,aes(x=GO_Term,y=number_of_gene,fill=Class))+geom_col(width=0.8)+
labs(y = "Number of Genes")+
geom_text(aes(x=GO_Term,y=number_of_gene,label=number_of_gene),size=2.5,hjust=0.5,vjust=-0.3)+
# 设置纵轴位置与刻度标签;
scale_y_continuous(limits = c(0, 80),
breaks = c(0,20, 40, 60,80),
label = c("0","20", "40", "60","80"),
expand = expansion(mult = c(0,0)),
position = "right")+
scale_x_discrete(expand = expansion(mult = c(0.2,0.05)))+
# 设置图表样式和图例位置;
theme_classic()+
theme(axis.title.x = element_blank(),
axis.text.x = element_blank(),
axis.line.x = element_blank(),
legend.position = "top",
legend.title = element_blank(),
legend.key.size = unit(3,"mm"),
plot.margin = margin(t = 0, r = 10, b = 0, l = 10, unit = "mm"))
# 自定义颜色;
p2 <- p1+scale_fill_manual(values = c("#FF99CC","#99CC00","#FF9900"))
p2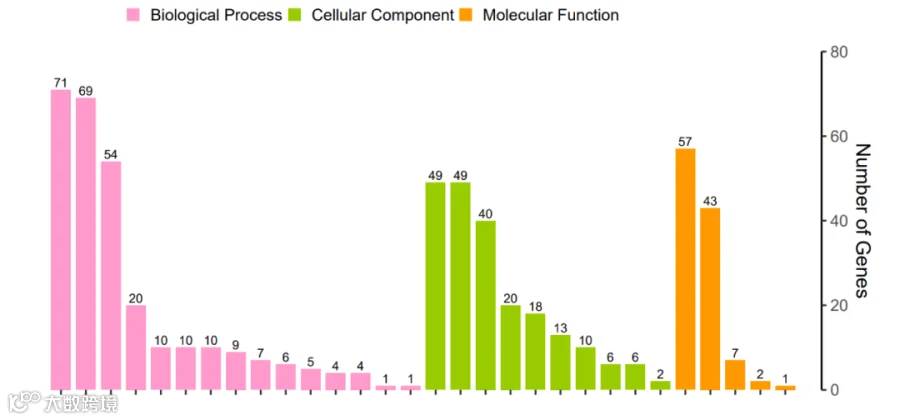
# 以新图表的方式单独绘制横轴标签,隐藏掉坐标轴;
f1 <- ggplot(df,aes(x=GO_Term,y=1))
f2 <- f1+geom_text(aes(x=GO_Term,y=0,label=GO_Term),size=3,angle=60,hjust=1,vjust=0.5)+
scale_y_continuous(expand = expansion(mult = c(0.2,0)),position = "right")+
scale_x_discrete(expand = expansion(mult = c(0.2,0.05)))+
theme_void()+
theme(axis.text = element_blank(),
axis.ticks = element_blank(),
axis.title = element_blank(),
plot.margin=margin(t = 1, r = 5, b = 1, l = 5, unit = "mm"))
# 创建线框路径的节点坐标数据框;
coord1 <- data.frame(x=c(0.4,-4.4,10.4,15.4),
y=c(0,-1,-1,0),
class=c(rep("BP",4)))
coord2 <- data.frame(x=c(0.6+15,-4.4+15,10.4+10,15.4+10),
y=c(0,-1,-1,0),
class=c(rep("CC",4)))
coord3 <- data.frame(x=c(0.5+15+10+0.1,-4.5+15+10+0.1,10.5+10+5+0.1,15.5+10+5+0.1),
y=c(0,-1,-1,0),
class=c(rep("MF",4)))
# 绘制线框;
f3 <- f2+geom_path(data=coord1,aes(x=x,y=y),color="#FF99CC",linewidth = 0.6)+
geom_path(data=coord2,aes(x=x,y=y),color="#99CC00",linewidth = 0.6)+
geom_path(data=coord3,aes(x=x,y=y),color="#FF9900",linewidth = 0.6)
f3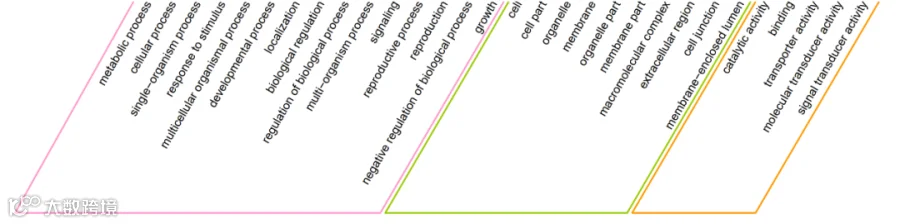
# 上下组合图表,完成图表绘制;
library(patchwork)
p2/f3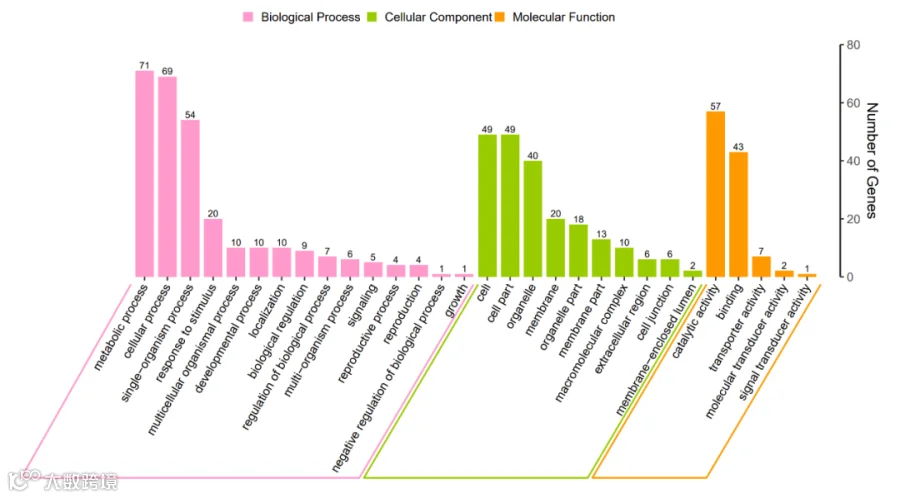
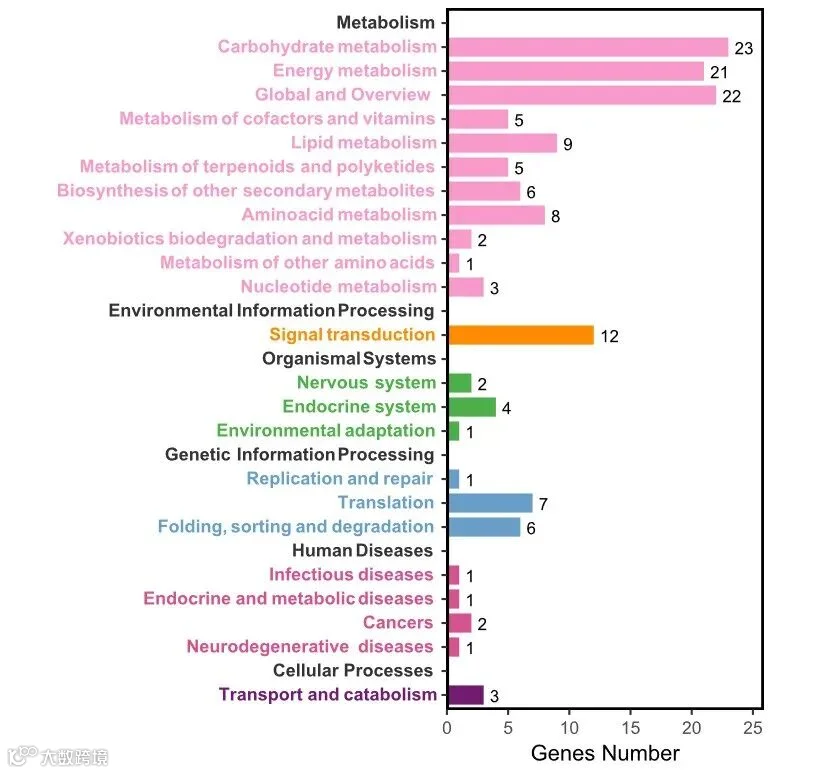
原教程链接:
关于环状条形图的绘制,推荐使用OmicShare的动态柱状图工具,无需考虑太多,只需按照示例文件整理数据即可。
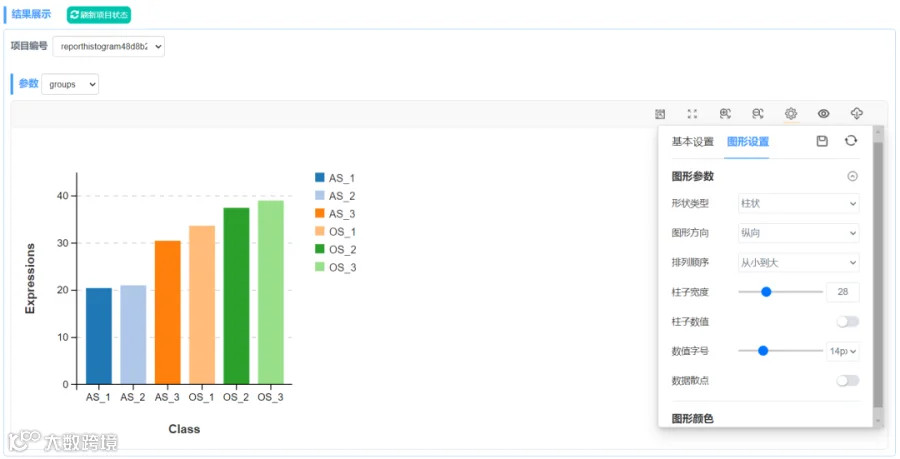
如果将形状类型设置为环状,则可以绘制文章中较为常见的环状条形图。
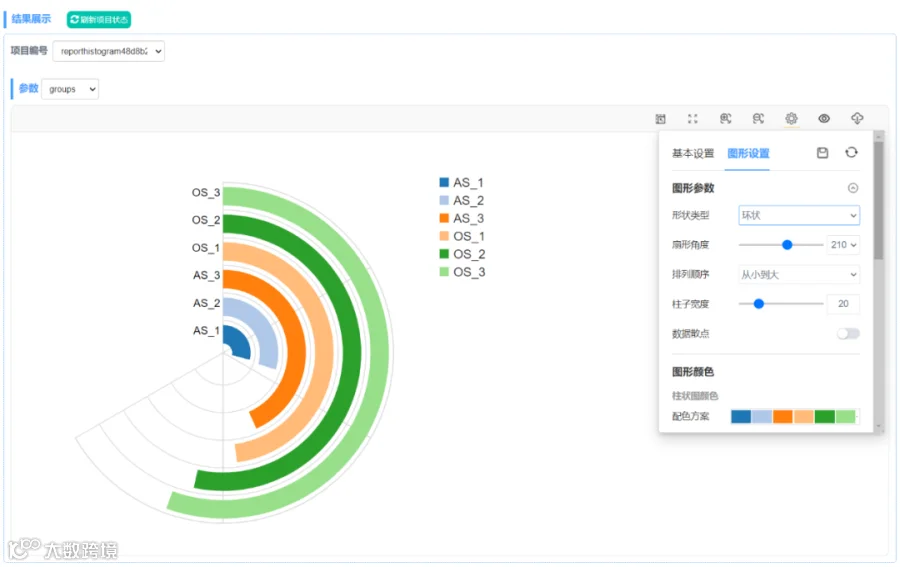
工具链接:
https://www.omicshare.com/tools/home/report/reporthistogram.html
原教程链接:
这种“相互嵌套”的柱状图,非常适合组合比较。一般来说,适合这种图表的数据,需要满足一组数据的数值低于另一组。
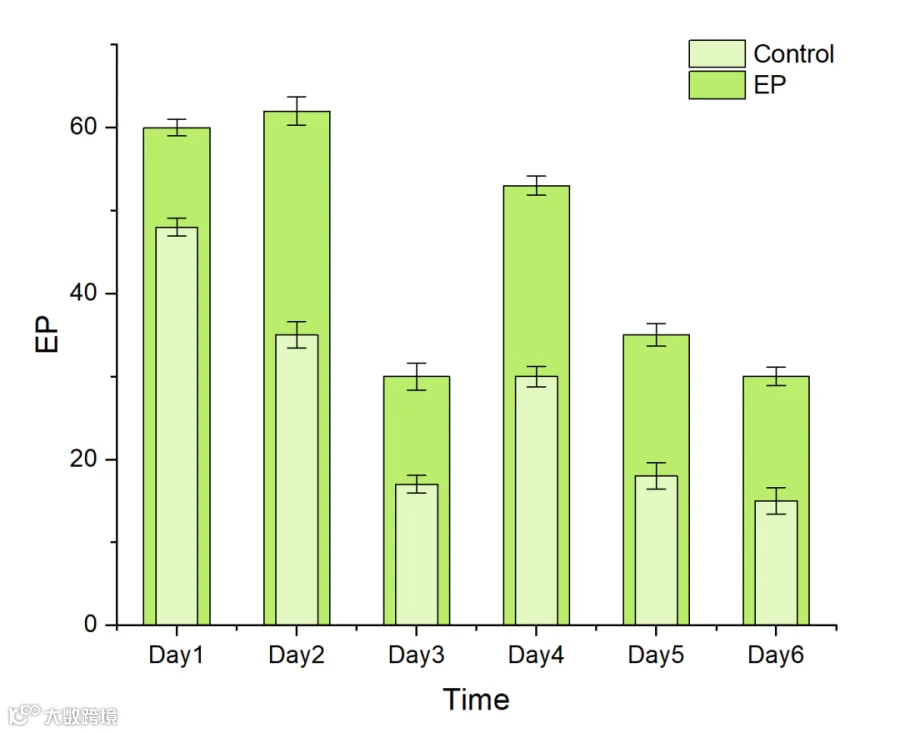
原教程链接:
使用Origin绘制分组柱状图,然后给Y轴加“break“,点击Y轴,右键选择Scale,弹出的窗口,在Break选项卡,Number of设为1,取消勾选Auto Position,自定义截断区域。绘制效果如下:
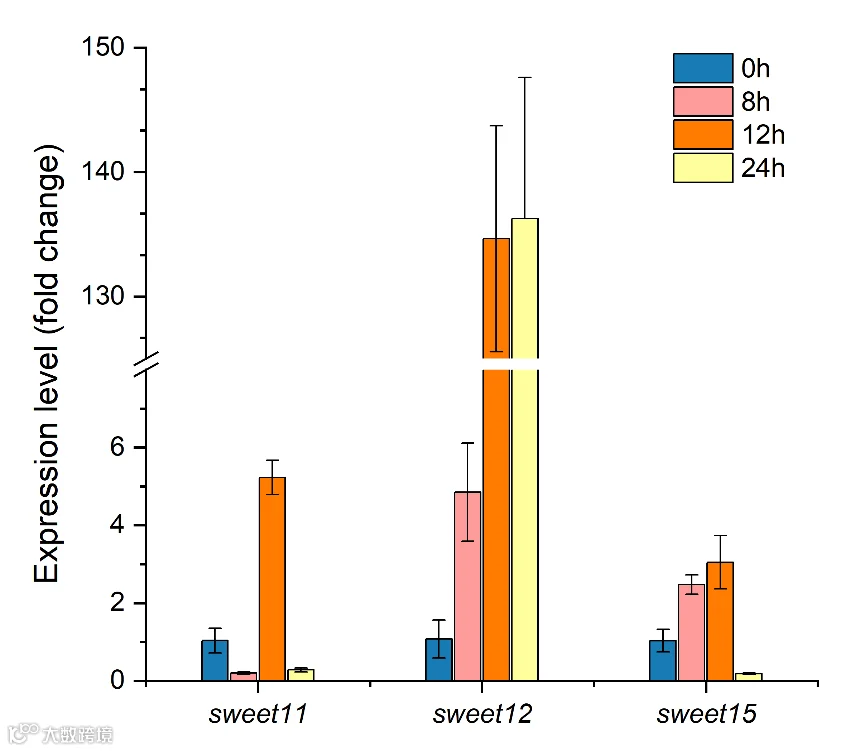
原教程链接:
Origin可绘制多种类型的3D图表,使用鼠标可以对视图“推拉摇移”,方便切换不同的视角进行观察。特别是绘制水晶质感的透视3D柱状图,特别好看。
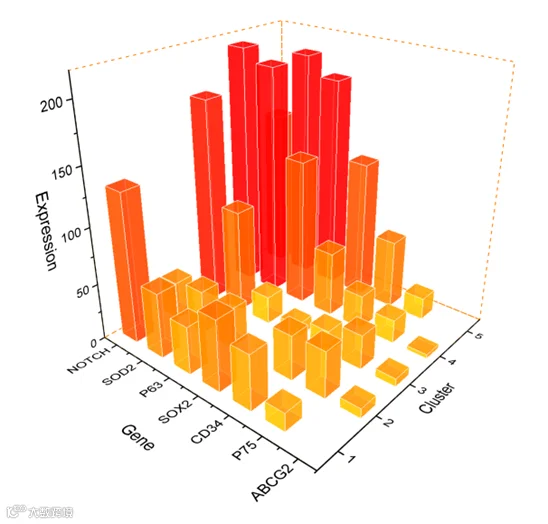
原教程链接:
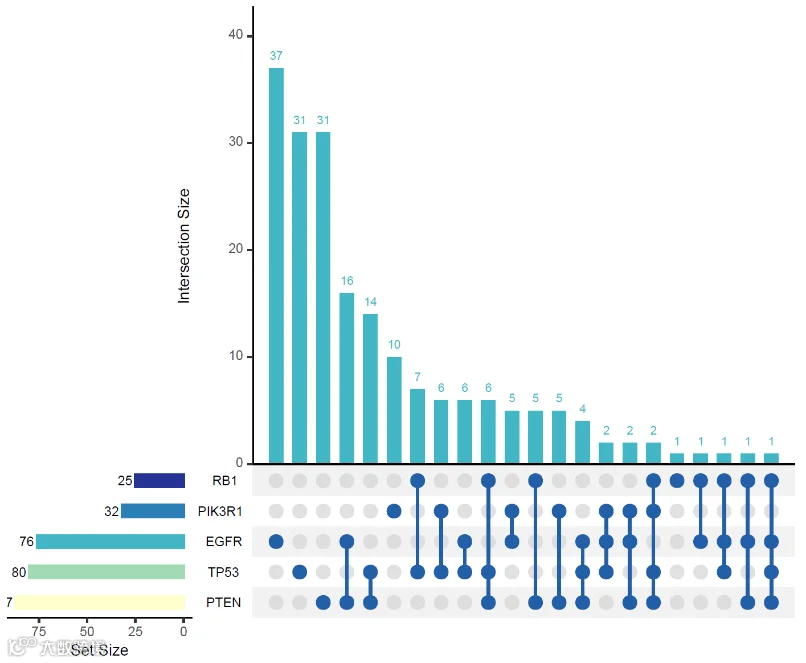
比较组数量过多时,可使用upset韦恩图进行展示,我们可以使用UpSetR包绘制该类图表。上方柱形图展示了集合特有或共有的交集元素数量,左方条形图展示各个集合中的元素总数,中心的交集矩阵中,单个颜色节点表示统计的特有元素信息,连线节点则表示统计的共有元素信息。
原教程链接:
好啦,本次的特色条形图绘制教程就分享到这里啦!
*未经许可,不得以任何方式复制或抄袭本篇文章之部分或全部内容。版权所有,侵权必究。

Omicsmaster
让每一分科研投入
都物超所值


