欢迎关注R语言数据分析指南
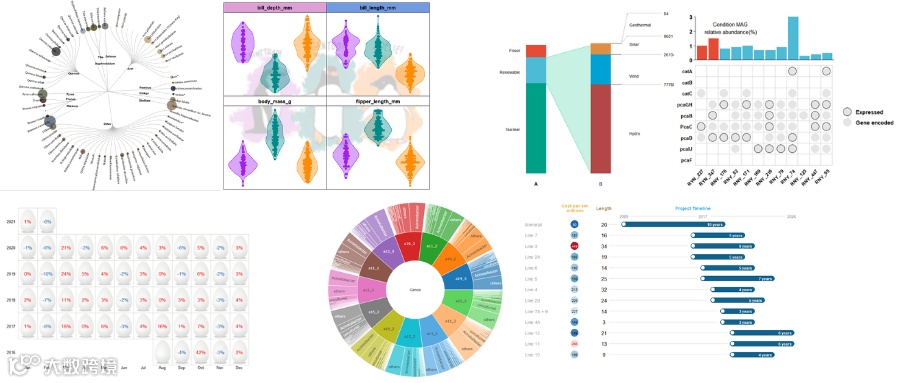
❝本节来介绍一个用于多个基因组的共线性和直系同源模式分析及可视化的R包「GENESPACE」,软件运行需要依赖其它软件如「OrthoFinder、MCScanX」等,分析环境配置好可以一站式完成数据的分析及可视化同时具有很高的自定性。小编下面进行部分的结果展示,软件安装等更多的详细内容请参考作者的官方文档。
❞
官方文档
❝https://github.com/jtlovell/GENESPACE
❞
(https://htmlpreview.github.io/?https://github.com/jtlovell/tutorials/blob/main/riparianGuide.html)
关注下方公众号下回更新不迷路
软件安装
if (!requireNamespace("devtools", quietly = TRUE))
install.packages("devtools")
devtools::install_github("jtlovell/GENESPACE")
library(GENESPACE)
定义软件路径
genomeRepo <- "~/path/to/store/rawGenomes"
wd <- "~/path/to/genespace/workingDirectory"
path2mcscanx <- "~/path/to/MCScanX/"
下载基因组数据
urls <- c(
human ="000/001/405/GCF_000001405.40_GRCh38.p14/GCF_000001405.40_GRCh38.p14_",
mouse = "000/001/635/GCF_000001635.27_GRCm39/GCF_000001635.27_GRCm39_",
platypus = "004/115/215/GCF_004115215.2_mOrnAna1.pri.v4/GCF_004115215.2_mOrnAna1.pri.v4_",
chicken = "016/699/485/GCF_016699485.2_bGalGal1.mat.broiler.GRCg7b/GCF_016699485.2_bGalGal1.mat.broiler.GRCg7b_",
sandLizard = "009/819/535/GCF_009819535.1_rLacAgi1.pri/GCF_009819535.1_rLacAgi1.pri_")
genomes2run <- names(urls)
urls <- file.path("https://ftp.ncbi.nlm.nih.gov/genomes/all/GCF", urls)
translatedCDS <- sprintf("%stranslated_cds.faa.gz", urls)
geneGff <- sprintf("%sgenomic.gff.gz", urls)
names(translatedCDS) <- genomes2run
names(geneGff) <- genomes2run
writeDirs <- file.path(genomeRepo, genomes2run)
names(writeDirs) <- genomes2run
for(i in genomes2run){
print(i)
if(!dir.exists(writeDirs[i]))
dir.create(writeDirs[i])
download.file(
url = geneGff[i],
destfile = file.path(writeDirs[i], basename(geneGff[i])))
download.file(
url = translatedCDS[i],
destfile = file.path(writeDirs[i], basename(translatedCDS[i])))
}
解析基因组注释文件
genomes2run <- c("human", "mouse", "platypus", "chicken", "sandLizard")
parsedPaths <- parse_annotations(
rawGenomeRepo = genomeRepo,
genomeDirs = genomes2run,
genomeIDs = genomes2run,
presets = "ncbi",
genespaceWd = wd)
软件运行
gpar <- init_genespace(
wd = wd,
ploidy = 1,
path2mcscanx = path2mcscanx)
out <- run_genespace(gpar, overwrite = T)
数据可视化
ripDat <- plot_riparian(
gsParam = out,
refGenome = "human",
forceRecalcBlocks = FALSE)

ripDat <- plot_riparian(
out,
refGenome = "human",
useOrder = FALSE,
useRegions = FALSE)

更改顺序
ripDat <- plot_riparian(
gsParam = out,
refGenome = "mouse",
genomeIDs = c("mouse", "human", "platypus", "chicken"),
forceRecalcBlocks = FALSE)

调整排序
ripDat <- plot_riparian(
gsParam = out,
#reorderBySynteny = FALSE,
syntenyWeight = 0,
refGenome = "human")

自定义标签
ggthemes <- ggplot2::theme(
panel.background = ggplot2::element_rect(fill = "white"))
customPal <- colorRampPalette(
c("darkorange", "skyblue", "darkblue", "purple", "darkred", "salmon"))
ripDat <- plot_riparian(
gsParam = out,
palette = customPal,
braidAlpha = .75,
chrFill = "lightgrey",
addThemes = ggthemes,
refGenome = "human")

部分区域高亮展示
roi <- data.frame(
genome = c("human", "chicken"),
chr = c("X", "Z"),
color = c("#FAAA1D", "#17B5C5"))
ripDat <- plot_riparian(
gsParam = out,
highlightBed = roi,
refGenome = "human",
genomeIDs = c("sandLizard", "chicken", "human", "mouse", "platypus"),
customRefChrOrder = c("X", 1:22))

❝可以看到内容还是非常的丰富,感兴趣的朋友可仔细阅读作者的官方文档;有需要学习R语言个性化数据可视化的朋友,欢迎到小编的「淘宝店铺」 「R语言数据分析指南」购买「2023年度会员文档」同步更新中「售价149元」,内容主要包括各种「高分论文的图表分析复现以及一些个性化图表的绘制」均包含数据+代码;按照往年数据小编年产出约在150+以上
❞
购买后微信发小编订单截图即邀请进新的会员交流群,小编的文档为按年售卖,只包含当年度的「除系列课程外」的文档,有需要往年文档的朋友也可下单购买,需要了解更多信息的朋友欢迎交流咨询。
淘宝扫一扫

2023会员群案例展示